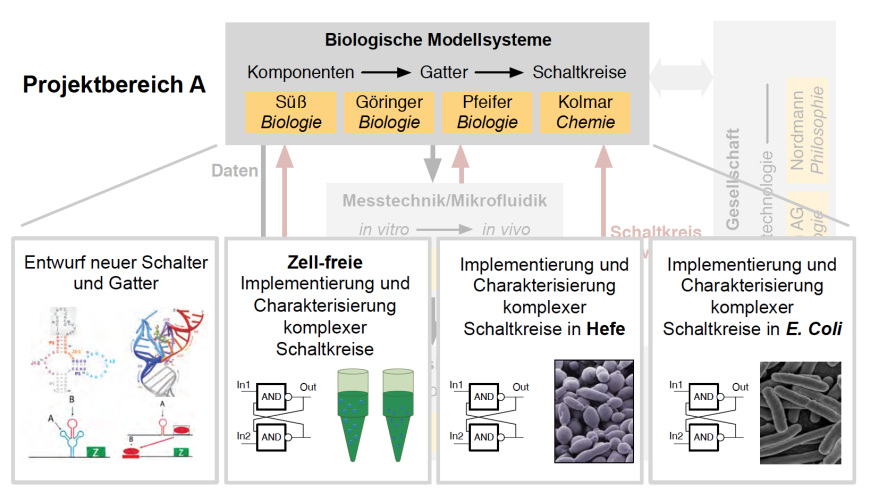
Diverse circuits with varying complexity should be implemented in E. coli and S. cerevisiae and the behavior of those circuits should be studied in context-dependent experiment.
Starting from simple inverting function the complexity of circuits will be raised until any desired number of input signals can be combined robustly. It will be necessary to optimize and even engineer completely now components. CompuGene has RNA-based regulatory elements on focus. Components and circuits will first be analyzed in vitro and than in the two different model organisms to demonstrate a common usability of the developed systems. So far, there is no need to leave the single cell organisms to higher complex. An important step is the precise analysis of dependency of genetic circuits from its conditions.
Project A-1
Characterization and optimization of biological components and simple connections towards circuit design
Project head: Beatrix Suess
Partial project A-1 aims at offering well characterized biological compounds and their logical connections for the synthesis of circuits. Those regulatory elements should be robust and universally usable coupled with selective switching behavior. The genomic context, external factors as well as cellular conditions should not have an influence on the elements. The collected data are forming the groundwork for the model building in partial project C and in the subsequent projects for circuit synthesis A-2 to A-4.
Project A-2
In vitro circuit implementation and adaptation of cell free system to in vivo conditions
Project head: H. Ulrich Göringer
In project A-2, individual logic gates or components of project A-1 will be connected in to modular circuits of higher complexity. Combinatorial or sequential circuits designs, for example, will thus be implemented. These can be combined to generate components such as memory, counters or adders, for example. A quantitative analysis of the multi modular systems will be achieved by measuring the structural consequence that results from joining individual components, as well as the performance of the implemented circuits with various in-vitro measurement systems.
Project A-3
Implementation of circuits in E. coli and Archaea
Project head: Felicitas Pfeifer
E. coli is the major model system used in the field of synthetic biology. Gained insights from project A-2 should therefore be validated in this organism. Bacteria-specific genetic circuits with diverse complexity should be build and characterized and in addition design proposals from project C should be considered, implemented and measured. Following aims have top priority: (1) Design and characterization of linear and backwards coupled circuits (shown in A-2, figure 5.4) consisting of multiple logical connections based on established and new compounds. (2) Comparison of circuits, which are build on transcriptional and/or translational regulatory elements. (3) Analysis of the specific context dependency in E. coli. (4) Transferability of circuit design onto other model organisms (Archaea)
Project A-4
Implementation of circuits in Saccharomyces cerevisiae
Project head: Harald Kolmar
Saccharomyces cerevisiae is not well studied compared to the model-organism E. coli, which plays the key role in synthetic biology. But the tremendous biotechnological relevance of S. cerevisiae leaves this research as a discrepancy. Therefore the development of routines to circuit synthesis is important. Partial project A-4 should generate methods for circuit synthesis in S. cerevisiae regarding eukaryotic-specific properties. A limiting factor for complex circuits is the number of available external inducers, therefore peptide-nucleic-acid (PNAs) should be considered as orthogonal regulatory component and implemented into circuit design.
Concrete goals of partial project A-4 are as follows:
(1) Are the compounds from A-1 and the insights regarding the circuit synthesis from A-2 and A-3 applicable to S. cerevisiae?
(2) Which are the organism-specific properties that should be considered for the circuit design and which yeast/eukaryotic-specific problems and context dependencies can arise?
(3) Are PNAs suitable for controlling circuits and how efficient is gene expression blocked?
(4) Is it applicable to use PNAs for logical connections? How does their switching properties behave and can they be used to build complex genetic circuits?